Distance Fields
- Overview
- Algorithms
- Visualizing distance
- Cylinder and torus distance
- Coordinate fields and Voronoi diagrams
- Procedural terrain
- References
Overview
Distance fields are useful in a variety of graphics applications, including antialiasing, ray marching, and texture synthesis. Sometimes they are computed analytically from functions, but often they are generated from voxelized meshes or 2D bitmaps. This article uses bitmaps for illustrative purposes.
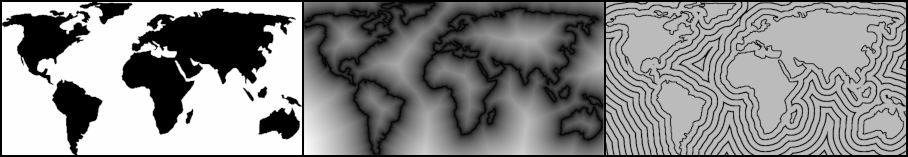
The EDT (Euclidean Distance Transform) can be defined as consuming a field of booleans and producing a field of scalars such that each value in the output is the distance to the nearest “true” cell in the input.
An example is shown in Figure 1b, but with a small twist; each distance value is squared (SEDT). By showing squared distance, we can can use integers everywhere in the diagram. The marching parabolas algorithm covered in the next section computes the SEDT, which is trivial to transform into the EDT.
Another useful concept is the signed distance field (SDF) which is the subtraction of the inverted EDT from the original EDT. This is depicted in Figure 1c. Note that negative values are inside the contour of the shape and positive values are outside. SDF’s play an important role in physics simulations and certain rendering techniques.
Algorithms
In the next two subsections I will describe two algorithms for generating distance fields from boolean fields:
- Marching Parabolas, a linear-time CPU-amenable algorithm.
- Min Erosion, a simple-to-implement GPU-amenable algorithm.
Marching Parabolas
This section is devoted to an algorithm for generating distance fields as described in 2012 by Felzenszwalb and Huttenlocher. It’s O(n) and astonishingly simple when compared to the alternatives that I came across.
The problem with transforming booleans into reals is that it prevents you from decomposing the 2D transform into two 1D transforms. If you’ve ever implemented an image filter you probably know that 2D convolution is best implemented using a horizontal pass followed by a vertical pass (or the reverse order).
So, the trick is to redefine the EDT so that you’re transforming reals to reals; this allows it to become a separable filter. Basically, you want each pixel in the input to have a scalar-valued coefficient of presence. We’ll do this by replacing all T values with zero, and all F values with infinity. The rationale for this will become apparent later in the article.
Figure 2 shows an example 10x10 image undergoing the series of transformations performed by the algorithm: horizontal distance transform, rotation, another horizontal transform, and finally another rotation.
Alternatively we could remove the rotations and change the second 1D transform into vertical pass, but I prefer thinking about the algorithm only in terms of horizontal passes. Sometimes this is actually more efficient, since rows have better cache coherency than columns.
The procedure depicted in Figure 2 is codified in Listing 1.
# Consume a 2D boolean field produce a 2D distance field.
def compute_edt(bool_field):
sedt = bool_field.where(0, ∞)
for row in len(sedt):
horizontal_pass(sedt[row])
transpose(sedt)
for row in len(sedt):
horizontal_pass(sedt[row])
transpose(sedt)
return sqrt(sedt)The transpose function is trivial to implement, so let’s focus on horizontal_pass. A key insight is that the 1D squared distance field is a set of samples from a series of overlapping quadratic parabolas. (Remember, we’re computing minimum squared distance.) This is made clear in Figure 3, which shows the second row of pixels from our example after the first pass.
Recall that we replaced all false values with infinity. In a sense, Figure 3 actually has ten parabolas: eight parabolas are high up at y=∞ (and therefore not visible), and two parabolas are at y=0. We can see now that a distance transform is really just a problem of finding the lower envelope in a series of parabolas.
So far we’ve only depicted parabolas at y=0, so let’s come up with a more interesting example, as would arise in the second 1D pass. See Figure 4.
Although Figure 4 has four parabolas, the lower envelope is composed of only three parabolas, and only two parabolas are actually sampled from. The following statements are now self-evident. By the way, the lowest point of a parabola is called its vertex.
- The 1D distance field is defined by the lower hull in a sequence of parabolas.
- Each parabola in the hull has the same shape but a unique position.
- Given a list of vertices and intersections in the lower hull, it is easy to compute the Y values by marching from left to right.
We can codify the process with the linear time algorithm in Listing 2.
def horizontal_pass(single_row):
hull_vertices = []
hull_intersections = []
find_hull_parabolas(single_row, hull_vertices, hull_intersections)
march_parabolas(single_row, hull_vertices, hull_intersections)Marching over the hull parabolas in order to populate the 1D distance values is fairly easy, see Listing 3. One gotcha is that multiple intersections can exist between two adjacent pixel centers, so we need an inner while loop. The inner loop usually has 0 or 1 iterations, so in practice this function is O(n).
def march_parabolas(single_row, hull_vertices, hull_intersections):
d = single_row
v = hull_vertices
z = hull_intersections
k = 0
for q in range(len(d)):
while z[k + 1].x < q:
k = k + 1
dx = q - v[k].x
d[q] = dx * dx + v[k].yNext let’s implement find_hull_parabolas, which actually solves an occlusion problem by removing parabolas that are completely above all the others.
This can be done by gradually building a list of parabolas from left to right and carefully tracking the intersection points. Each parabola-to-parabola intersection can be computed with simple algebra. The procedure for this is shown in Listing 4.
def find_hull_parabolas(single_row, hull_vertices, hull_intersections):
d = single_row
v = hull_vertices
z = hull_intersections
k = 0
v[0].x = 0
z[0].x = -INF
z[1].x = +INF
for i in range(1, len(d)):
q = (i, d[i])
p = v[k]
s = intersect_parabolas(p, q)
while s.x <= z[k].x:
k = k - 1
p = v[k]
s = intersect_parabolas(p, q)
k = k + 1
v[k] = q
z[k].x = s.x
z[k + 1].x = +INF
# Find intersection between parabolas at the given vertices.
def intersect_parabolas(p, q):
x = ((q.y + q.x*q.x) - (p.y + p.x*p.x)) / (2*q.x - 2*p.x)
return x, _Note that the intersection computation does not bother returning a valid Y value, it is not needed.
When a new parabola is added it to the hull, the algorithm checks if the previously-added parabola is “above” the new parabola, in which case it gets removed from the hull. This process is depicted in the following animation (web only), which uses the same source data that was used to generate Figure 4.
To summarize, we can create a 1D distance field by executing a series of linear-time procedures:
- Interpret each row in the source image as a list of parabolas at various heights.
- Determine the set of parabolas that form the outer hull.
- March through the parabolas in the hull, computing the Y value at each pixel center.
One nice property of this algorithm is that the computed distance field need not have the same resolution as the source data, since the final values are computed by a sampling from a list of parabolic functions.
To see a full implementation of the marching parabolas algorithm, see the references section.
Min Erosion
The marching parabolas algorithm can be implemented using any general purpose programming language, but some situations call for an algorithm that is easy to implement using a graphics API such as OpenGL, WebGL, Vulkan, or Metal.
One way of doing this involves a series of image processing passes, where each pass is a full-screen quad or triangle. This works by applying a series of horizontal passes followed by a series of vertical image processing passes, as shown below. Note that this once again computes squared distance, and that the seed image is once again composed of values 0 and “infinity” (where infinity is represented by the F hexadecimal digit).
The top row depicts a series of horizontal passes, the bottom row shows the vertical passes. Pixels outlined in yellow are modified while all other pixels are discarded. The letters A and B refer to a pair of ping-pong buffers since GPU’s typically do not allow sampling from arbitrary locations in the render target. β is an odd integer associated with pass and is increased by 2 after each pass.
In each pass, every pixel is compared to two neighboring values that have been augmented by adding β. If either of the augmented neighbor values are less than the value at the current pixel, the current pixel’s value is overwritten with the augmented value.
Note that a given pixel might be modified more than once! For example, the F in the lower-left corner changes to a 9 before settling to its final value of 8.
Thus, each sequence of passes terminates only when there are no further modifications that can be made (i.e. when all pixels are discarded). This can be detected using a query such as GL_ANY_SAMPLES_PASSED. Alternatively, a reasonably accurate approximation to the distance field can be computed by simply terminating after a predetermined number of passes.
Some example GLSL shader code for the above method is available here. This code produces an RGB image that has squared distance in the blue channel and the closest point coordinate stored in the red-green channels. We will discuss closest point coordinates later in the post.
For a more efficient GPU-amenable method, see also jump flooding by Rong and Tan. Their method generates a closest point coordinate field from which a distance field can be derived.
Also note that distance fields can be composed together by using a special blending operation that takes the minimum of the source and destination color. In OpenGL, this is enabled by passing GL_MIN into glBlendEquation. For example, one (impractical) way of generating a distance field from a small point cloud would be to render a large quad centered at each point, where the fragment shader computes a brightness proportional to the distance from the quad’s center.
Visualizing distance
Let’s set aside generation of distance fields and instead discuss methods of depicting them.
The easiest way to visualize a SDF is to take the absolute value of each pixel, then normalize the values such that the maximum distance becomes white and the minimum distance becomes black. This is depicted in the middle panel in Figure 5.

Another strategy is to draw contour lines, shown in the right panel. This can be done by blackening out distance values that fall within a series of equally-spaced ranges. Here’s an example implementation using numpy:
for h in range(0, 1000, 100):
this_contour = np.logical_and(sdf >= h-1, sdf <= h+1)
all_contours = np.logical_or(all_contours, this_contour)Cylinder and torus distance
By stacking copies of the source image on either side, then extracting the middle portion from the resulting distance field, it can be made tileable. This could be useful if you know that your distance field will be wrapping a cylinder (or a sphere, using a lat-long projection).
Figure 6 shows a tileable EDT. Note that the contour lines to the far west of South America are quite different from the non-tilable version in Figure 5.
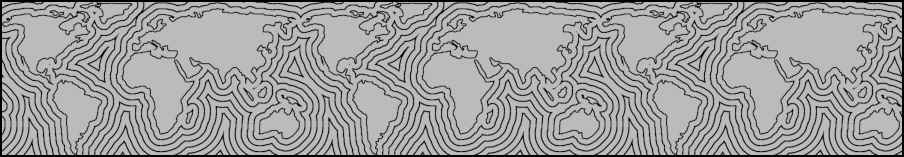
Toroidal wrapping can be achieved by creating an EDT from 3x3 tiling of the source image, then extracting the middle tile from the result.
Coordinate fields and Voronoi diagrams
The closest point transform (CPT) is related to the EDT. It consumes a field of booleans and produces a field of coordinates, where each coordinate points to the nearest T in the input field.
The CPT is easy to transform to the EDT: for each pixel, simply compute the distance between that pixel and the pixel that the CPT points to.
The CPT is also easy to transform to a Voronoi diagram: for each pixel, replace it with the color of the pixel that the CPT points to. This is illustrated in Figure 7. This is actually a generalized Voronoi diagram because the source image is not composed of discrete points.
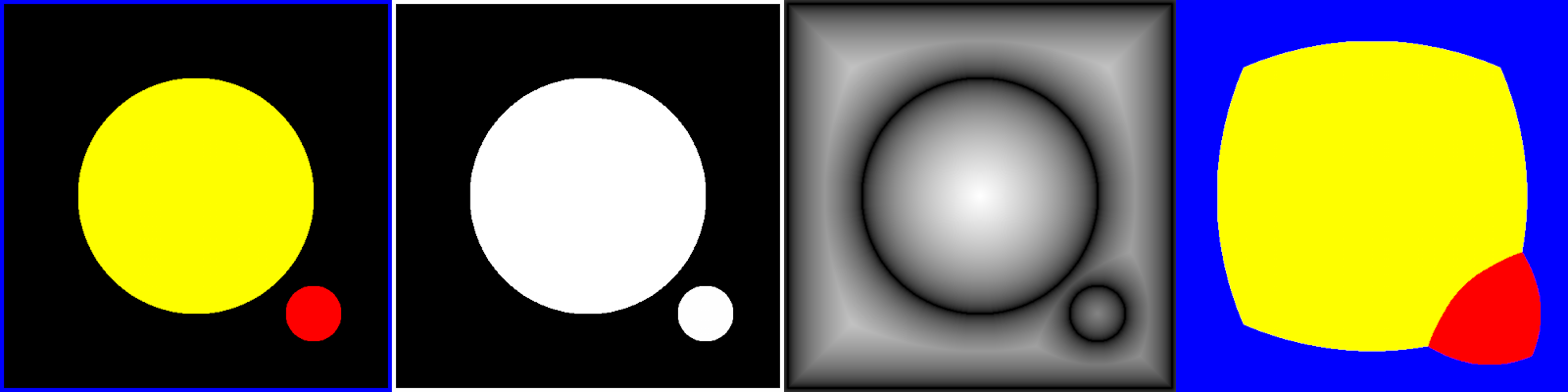

To generate the CPT, the march_parabolas procedure can be modified to store the column index of each parabola vertex. See Listing 5, which differs from Listing 3 in that it adds an indices argument that gets populated with the relevant X coordinates.
def march_parabolas(single_row, hull_vertices, hull_intersections, indices):
d = single_row
v = hull_vertices
z = hull_intersections
k = 0
for q in range(len(d)):
while z[k + 1].x < q:
k = k + 1
dx = q - v[k].x
d[q] = dx * dx + v[k].y
indices[q] = v[k].xThe first call to horizontal_pass produces X coordinates, and the second call to horizontal_pass produces Y coordinates. Afterwards, the X coordinates need to be dereferenced to obtain the final coordinate field. This procedure is outlined in Listing 6, which can be compared to Listing 1.
def compute_cpt(bool_field):
sedt = bool_field.where(0, ∞)
xcoords = empty 2D field of scalars
ycoords = empty 2D field of scalars
for row in len(sedt):
horizontal_pass(sedt[row], xcoords)
transpose(sedt)
for row in len(sedt):
horizontal_pass(sedt[row], ycoords)
# Deference the X coordinates to produce the final CPT.
cpt = empty 2D field of coordinates
for j in height:
for i in width:
x = xcoords(i, j)
y = ycoords(i, j)
cpt(i, j) = (cpt(i, y).x, y)
return cptProcedural terrain
Distance fields can be used to generate reasonable height maps for procedurally-generated terrain. Mountain chains tend to follow the “spine” of the shape. To mitigate the artificial look, gradient noise can be used to adjust the generated elevation data.
Figure 8 shows one possible way of generating a source mask for a landmass. From left to right: falloff function, four octaves of gradient noise multiplied with the falloff, warping, masking.
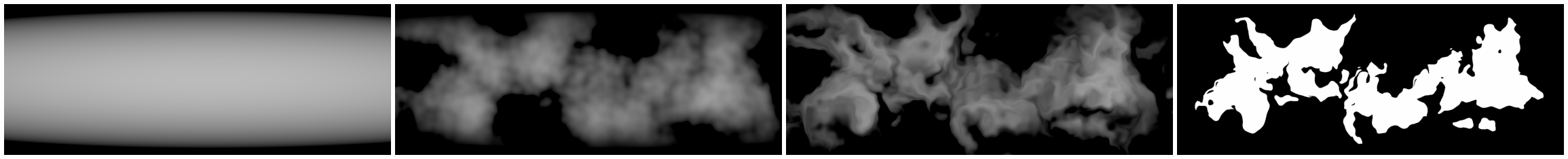
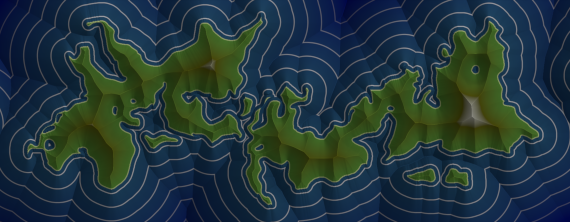
Figure 9 shows a rendering of the SDF computed from the above mask. This was rendered by multiplying three layers: diffuse lighting, ambient occlusion, and a color gradient.
Another interesting look can be achieved by quantizing the distance field using a step function. The boundaries between the discrete values suggest contour lines. The image in Figure 10 was generated by processing the EDT with the numpy digitize function.
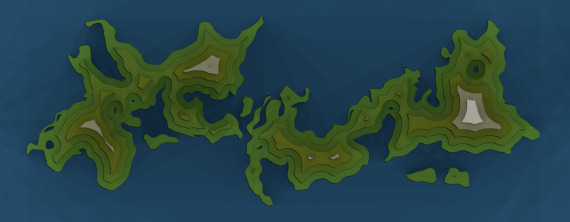
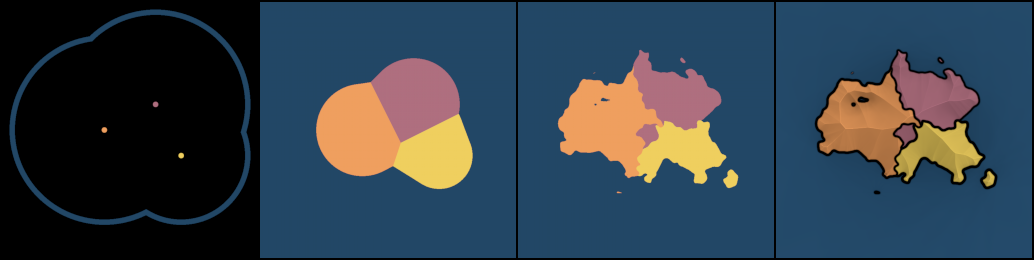
To generate random political regions, coordinate fields from the previous section can be used in combination with noise-based warping; see figure 11.
References
Inigo Quilez has a wealth of information about analytic distance fields at his blog, and these are some of my favorite references:
Implementations of the “parabolas” algorithm are available from a few different sources:
- Felzenszwalb page at Brown University (C++06)
- Giorgio Marcias github project (C++11)
- snowy (Python 3)
- heman (C99)
- nile (nim)
Another interesting library is DGtal, which can generate N-dimensional data and even has a reverse distance transform.
For generating distance fields from bitmaps on the GPU, check out the following paper from Rong and Tan.
This is a monograph on distance fields, written by Philip Rideout in 2018. If you’d like to print this out, you can download a pdf file.